March 2018
A modern era of targeted crop breeding is upon us. Traditional plant breeding aims to change traits/genes in a specific crop to obtain the desired characteristics in the offspring. This targeted trait is often linked to a specific change (mutation) in the parental plants’ genetic code (DNA), which the breeder then attempts to develop progeny from, containing target market characteristics.
Though this process seems relatively straight forward it is not, since some traits are ‘hidden’ (in the form of recessive genes), while other traits are transferred to the progeny in large groups (linked) that may include undesirable traits as well (linkage drag).
Unwanted traits are also randomly transferred to the progeny, which means as the number of desired traits increases, the number of progeny required to obtain an individual with all the desired traits and development cost, increases dramatically. This numbers game becomes even more complicated when breeding with grain crops. This is especially the case for wheat, with its three large complex genomes, having multiple copies of a single gene that originated from the ancestry donor backgrounds.
The breeding process always aims to produce cultivars faster and therefore currently uses tools such as molecular selection (marker-assisted breeding), embryo rescue and double haploid generation and speed breeding to accomplish this.
This ultimately results, after many years (eight to twelve years) of breeding and selection cycles (including traditional trait screening and molecular selection), in the release of higher yielding and adapted cultivars.
New plant breeding technologies targets traits better
However, what if a breeder could actually ‘target’ a desired trait with precision, thereby transferring only the new desired trait into an elite line without other unwanted characteristics? Well, some of the new breeding technologies in the breeder’s toolbox will now allow just that: The ability to transfer a specific trait by targeting the specific genetic code or gene region responsible for it.
These new plant breeding tools include a wide variety of technologies, ranging from directed nucleases for targeted mutagenesis to technologies that transfer the trait of interest but does not result in permanent DNA changes.
The tools in the new breeding toolbox that are really making a huge impact are those belonging to the directed nucleases group. Nucleases are enzymes that can cut DNA. Some of these nucleases recognise and cut only specific DNA sequences (e.g. meganucleases), while others use engineered proteins to target specific DNA for cleaving. The usefulness of meganucleases are limited since they can only target and cut at their specific DNA recognition sequences, which will very rarely be within the target
trait region desired by the breeder. These technologies require expensive, time-consuming protein engineering skills by experienced individuals. All these directed nucleases have been around for a number of years, but not widely adopted due to their limitations. This all changed with the discovery of the CRISPR/Cas system – a system awarded the Science Discovery of the Year in 2015.
CRISPR/Cas: Get familiar with it
CRISPR/Cas9 (clustered regularly interspaced short palindromic repeat)-Cas9 is a multipurpose system for targeted genetic engineering that uses a ‘guide molecule’ (guide RNA) to direct a DNA nuclease (Cas9 or other similar endonuclease) to a specific target site where it cuts the DNA in a specific manner.
The guide can easily and quickly be changed and synthesised as DNA (or RNA) using current technologies, making this system not only cheaper, but also faster to implement, with less expertise required. The usage of the system is restricted by a Cas nuclease specific protospacer adjacent motif (PAM) – a short, Cas enzyme specific sequence, required to be in the target DNA for successful binding and cutting (Figure 1).
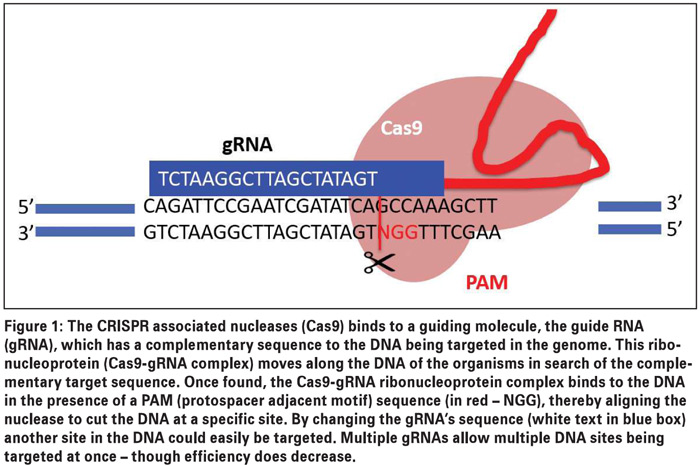
This PAM sequence differs between different Cas nucleases, thereby increasing possible cutting sites and allowing easier targeting of a different desired DNA sequence. The availability of complete genome sequence, target gene sequence/mutation and appropriate in vitro (tissue culture) delivery system, are some of the major limitations of the system currently.
How does cutting DNA help breeding?
But how can just cutting DNA, result in a targeted trait introduction (mutation)? This is where the plant’s (actually any organism) natural DNA repair mechanisms come into play. During day-to-day activities or through environmental influences, DNA strands may naturally break. Therefore, each cell has the capacity to fix such breaks using one of two processes: Non-homologous end joining (NHEJ) or homologous repair (HR) (Figure 2).
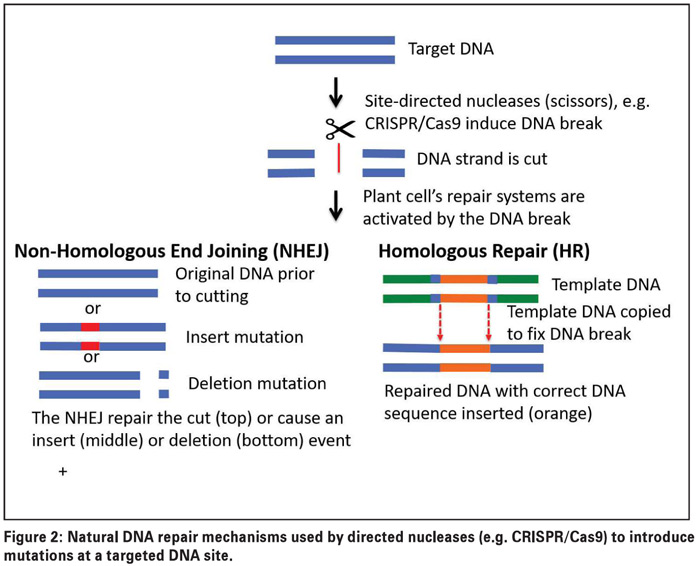
The NHEJ system can fix the break by just joining the two DNA strands again, or through an error-prone system that naturally results in small DNA mutations (changes) by either adding or removing small DNA pieces. The HR system uses an existing copy of the DNA as a template to precisely correct an error.
Scientists found that when a double stranded break is introduced into a DNA strand, one of these two mechanisms would be activated naturally to repair the break. Most often the NHEJ repair system would be activated, to either join the DNA strands again – thus reforming the original DNA sequence – or introducing a mutation by deleting or inserting DNA pieces.
A non-mutational repair results in rebuilding of the original targeted site, thus allowing the CRISPR/Cas system or other-directed nucleases to cut it again. A mutation (DNA insert or deletion), changes this target site, making it non-recognisable to the nucleases. HR is used for more precise changes (mutations) when a ‘correct DNA template’ is provided with the site directed nucleases from which the cell can ‘copy’ from.
This provides the breeder with the tools to ‘knock-out’ (switch a gene/trait off) a gene or to even change a single DNA base, thereby allowing the introduction of a new trait or edited target gene. This is very similar, if not identical, to traditional mutation breeding in the past, but it is much more targeted for a specific trait of interest. ‘Knocking’ a trait out is currently considered easier than introducing one. This tool is seen as vital technology going forward which will have the largest impact during early generations of the crop development process.
What it means for the producer or end user
This technology will be used with increasing frequency in the breeding process in the future to generate new improved higher-yielding crop cultivars in half the time. It will depend on which private companies/research institutions are willing to invest in it for the long-term and how it will be regulated.
This tool has already been used successfully internationally in the development of improved cultivars in crops like maize, wheat, rice, tomato, soybean, with target traits relating to nutritional quality, drought tolerance, disease and herbicide resistance.
It has been successfully used on a number of important crops to introduce herbicide resistance to chlorsulfuron by means of single DNA base change. Indeed, such herbicide resistance has already been introduced into maize, tomato and rice using the CRISPR technology.
The technology is currently being used in livestock research to promote animal health and well-being, with CRISPR/Cas aided dehorning already shown to work in dairy cattle.
Safety first
With all these new plant breeding tools and activities happening around the world, one would expect to see large numbers of improved crops being released already. However, this is not the case. Why?
The success and speed at which the new tools were developed and released caught the regulatory bodies across the world off guard. Many countries are just not sure how the technologies should be regulated or even if they should for certain usages. What is especially concerning is that literally anyone can experiment with the CRISPR/Cas system.
This is a real possibility as many CRISPR/Cas tutorial videos and readymade system kits are readily available for purchase online. Someone can experiment with the CRISPR/Cas system in their backyard or ‘garage lab’ for good or bad. This is further complicated since some of the technologies could also be used for transgene events, i.e. generating genetically modified organisms (GMOs) – which are indeed regulated in many countries, including South Africa.
Food for thought
However, how do you regulate a mutation that was induced using the existing natural mechanisms (HR or NHEJ) within the organism – either directed using the new breeding tools, or induced using non-regulated, random mutagenesis technologies or even occurring naturally – especially if you cannot determine the route taken by looking at the end product only. The USA is taking the lead, with a CRISPR/Cas9 edited, non-browning longer shelf life mushroom recently approved for limited release.
The EU recently had a workgroup looking into the regulatory requirements of each technology and it has made some recommendations already.
South Africa is also proactive in this regard with the Department of Science and Technology funding the Academy of Science of South Africa’s (ASSAF) investigation into these tools and their regulation. Their findings and reviews from other countries’ current regulations were last year published in a report on new breeding technologies (available from their webpage). They concluded that these technologies could make use of the current regulatory framework of the GMO act, since the act ensures that any transgene events would be adequately regulated as per SA law, while non-GM events would exit from the process earlier.
South Africa, will however, most likely keep in line with the regulations of their key trade partners such as the EU, to ensure compliance and continued trade.
What’s around the corner?
Targeted breeding technologies, like CRISPR/Cas, are here to stay and will increasingly play a bigger role in our daily lives. Not only, helping with the development of new high yielding cultivars or livestock with superior characteristics and quality in far shorter time, but also allow us to address human disease at the molecular level.
 In future, more medical treatments will rely solely on the CRISPR/Cas technology to target cancers (leukaemia) and inheritable diseases than the current limited cases in the USA and UK. Indeed, an exciting time to be a breeder and a potentially brighter future for our producers.
In future, more medical treatments will rely solely on the CRISPR/Cas technology to target cancers (leukaemia) and inheritable diseases than the current limited cases in the USA and UK. Indeed, an exciting time to be a breeder and a potentially brighter future for our producers.

Publication: March 2018
Section: Focus on